3 Do-it-yourself with R
In this book, we introduce various methods for instance-level and dataset-level exploration and explanation of predictive models. In each chapter, there is a section with code snippets for R and Python that shows how to use a particular method. In this chapter, we provide a short description of the steps that are needed to set-up the R environment with the required libraries.
3.1 What to install?
Obviously, the R software (R Core Team 2018) is needed. It is always a good idea to use the newest version. At least R in version 3.6 is recommended. It can be downloaded from the CRAN website https://cran.r-project.org/.
A good editor makes working with R much easier. There is plenty of choices, but, especially for beginners, it is worth considering the RStudio editor, an open-source and enterprise-ready tool for R. It can be downloaded from https://www.rstudio.com/.
Once R and the editor are available, the required packages should be installed.
The most important one is the DALEX package in version 1.0 or newer. It is the entry point to solutions introduced in this book. The package can be installed by executing the following command from the R command line:
install.packages("DALEX")Installation of DALEX will automatically take care about installation of other requirements (packages required by it), like the ggplot2 package for data visualization, ingredients and iBreakDown with specific methods for model exploration.
3.2 How to work with DALEX?
To conduct model exploration with DALEX, first, a model has to be created. Then the model has got to be prepared for exploration.
There are many packages in R that can be used to construct a model. Some packages are algorithm-specific, like randomForest for random-forest classification and regression models (Liaw and Wiener 2002), gbm for generalized boosted regression models (Ridgeway 2017), extensions for generalized linear models (Harrell Jr 2018), and many others. There is also a number of packages that can be used for constructing models with different algorithm These include the h2o package (LeDell et al. 2019), caret (Jed Wing et al. 2016) and its successor parsnip (Kuhn and Vaughan 2019), a very powerful and extensible framework mlr (Bischl et al. 2016), or keras that is a wrapper to Python library with the same name (Allaire and Chollet 2019).
While it is great to have such a large choice of tools for constructing models, the disadvantage is that different packages have different interfaces and different arguments. Moreover, model-objects created with different packages may have different internal structures. The main goal of the DALEX package is to create a level of abstraction around a model that makes it easier to explore and explain the model.
Function DALEX::explain is THE function for model wrapping. There is only one argument that is required by the function; it is model, which is used to specify the model-object with the fitted form of the model. However, the function allows additional arguments that extend its functionalities. They will be discussed in Section 5.2.6.
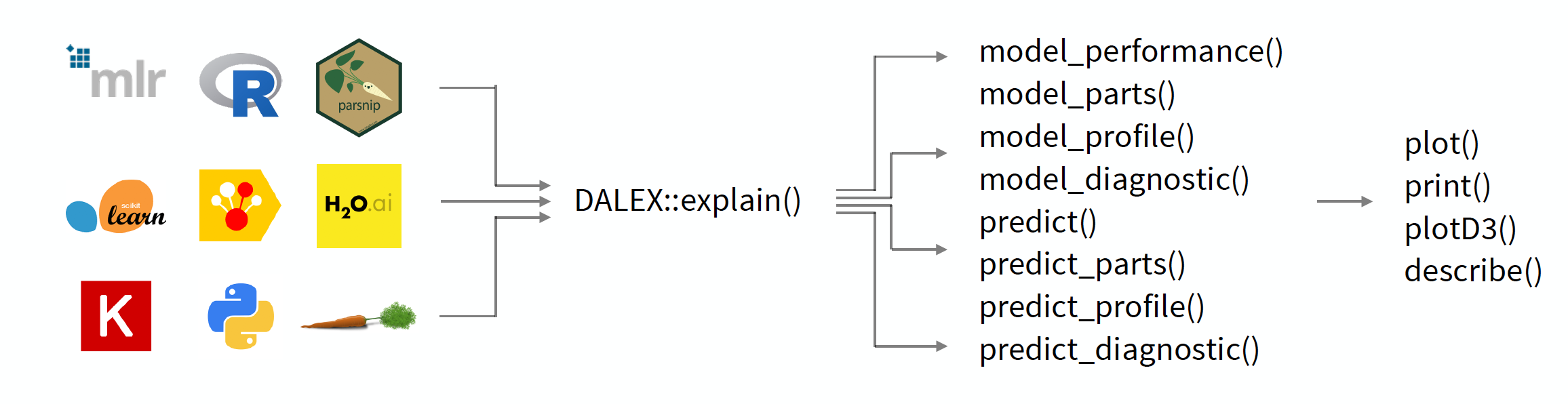
Figure 3.1: The DALEX package creates a layer of abstraction around the models, allowing you to work with different models in a uniform way. The key function is the explain function, which wraps any model into a uniform interface. Then other functions from DALEX package can be used on the resulting object to explore the model.
3.3 How to work with archivist?
As we will focus on the exploration of predictive models, we prefer not to waste space nor time on replication of the code necessary for model development. This is where the archivist packages help.
The archivist package (Biecek and Kosinski 2017) is designed to store, share, and manage R objects. We will use it to easily access pretrained R models and precalculated explainers. To install the package, the following command should be executed in the R command line:
install.packages("archivist")Once the package has been installed, function aread() can be used to retrieve R objects from any remote repository. For this book, we use a GitHub repository models hosted at https://github.com/pbiecek/models. For instance, to download a model with the md5 hash ceb40, the following command has to be executed:
Since the md5 hash ceb40 uniquely defines the model, referring to the repository object results in using exactly the same model and the same explanations. Thus, in the subsequent chapters, pre-constructed models will be accessed with archivist hooks. In the following sections, we will also use archivist hooks when referring to datasets.
References
Allaire, JJ, and François Chollet. 2019. Keras: R Interface to ’Keras’. https://CRAN.R-project.org/package=keras.
Biecek, Przemyslaw, and Marcin Kosinski. 2017. “archivist: An R Package for Managing, Recording and Restoring Data Analysis Results.” Journal of Statistical Software 82 (11): 1–28. https://doi.org/10.18637/jss.v082.i11.
Bischl, Bernd, Michel Lang, Lars Kotthoff, Julia Schiffner, Jakob Richter, Erich Studerus, Giuseppe Casalicchio, and Zachary M. Jones. 2016. “mlr: Machine Learning in R.” Journal of Machine Learning Research 17 (170): 1–5. http://jmlr.org/papers/v17/15-066.html.
Harrell Jr, Frank E. 2018. Rms: Regression Modeling Strategies. https://CRAN.R-project.org/package=rms.
Jed Wing, Max Kuhn. Contributions from, Steve Weston, Andre Williams, Chris Keefer, Allan Engelhardt, Tony Cooper, Zachary Mayer, et al. 2016. Caret: Classification and Regression Training. https://CRAN.R-project.org/package=caret.
Kuhn, Max, and Davis Vaughan. 2019. Parsnip: A Common Api to Modeling and Analysis Functions. https://CRAN.R-project.org/package=parsnip.
LeDell, Erin, Navdeep Gill, Spencer Aiello, Anqi Fu, Arno Candel, Cliff Click, Tom Kraljevic, et al. 2019. H2o: R Interface for ’H2o’. https://CRAN.R-project.org/package=h2o.
Liaw, Andy, and Matthew Wiener. 2002. “Classification and Regression by randomForest.” R News 2 (3): 18–22. http://CRAN.R-project.org/doc/Rnews/.
R Core Team. 2018. R: A Language and Environment for Statistical Computing. Vienna, Austria: R Foundation for Statistical Computing. https://www.R-project.org/.
Ridgeway, Greg. 2017. Gbm: Generalized Boosted Regression Models. https://CRAN.R-project.org/package=gbm.